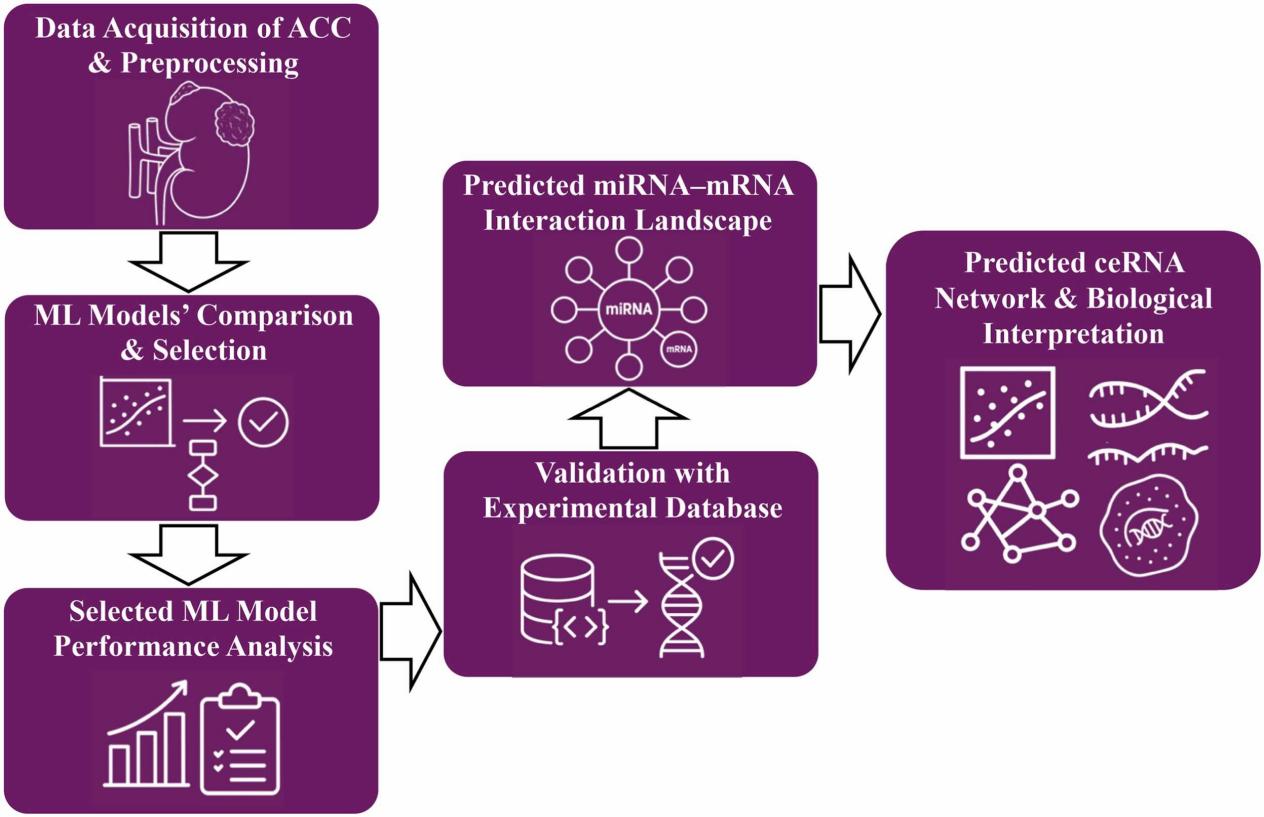
A new study published in Intelligent Oncology presents a machine learning-driven integrative framework to systematically decode the miRNA-mRNA ceRNA regulatory network in adrenocortical carcinoma (ACC), offering fresh insights into biomarker discovery and target prioritization for this rare malignancy.
Background
Adrenocortical carcinoma is a rare but highly aggressive endocrine malignancy, with an annual incidence of only 0.7–2 cases per million. However, due to its high metastatic potential and resistance to conventional therapy, the 5-year survival rate for patients with advanced disease falls below 40%. There is an urgent clinical need for reliable molecular biomarkers and novel therapeutic targets.
Within competing endogenous RNA (ceRNA) networks, microRNAs (miRNAs) act as key regulatory hubs by interacting with messenger RNAs (mRNAs) to influence tumor development. Traditional correlation-based methods struggle to capture nonlinear, context-specific regulatory relationships, highlighting the need for more powerful computational models.
Study Overview
To address this challenge, the researcher integrated RNA-Seq and miRNA-Seq data from TCGA-ACC (78 tumor samples) and GTEx (250 normal adrenal samples). Five regression models [Random Forest (RF), XGBoost, LightGBM, Gradient Boosting Regressor, and Linear Regression] were systematically benchmarked, with RF emerging as the best-performing algorithm.
Model performance was impressive:
Normal tissue: R2=0.9467, MAE = 0.0116
Tumor tissue: R2=0.9045, MAE = 0.0242
The study further employed SHAP values for model interpretability, identifying tumor-specific miRNAs (e.g., hsa-mir-205, hsa-mir-4772) that exert a strong influence on the regulatory network.
Key Findings
The study revealed several important results:
Topological rewiring: Compared to normal tissue, ACC tumor tissue exhibited a pronounced loss of regulatory hub connectivity, with a sparser and more centralized network architecture.
Novel high-confidence interactions: 64 tumor-specific, high-scoring miRNA-mRNA pairs were identified, including eight particularly prominent pairs involving genes such as MMP2, COL6A3, IL24, and BMP3, which are implicated in tumor invasion and immune signaling.
Overlap with reference databases: Predicted interactions showed good concordance with established databases such as miRTarBase and TargetScan (e.g., 36/100 top tumor pairs were supported by both databases), reinforcing the biological credibility of the model.
Clinical and Research Implications
The author notes that this machine learning-driven approach has the potential to:
Provide a systematically prioritized resource of ceRNA interactions to guide future experimental validation;
Reveal tumor-specific regulatory hubs (e.g., hsa-mir-205, SCNN1A, EFNA1) as promising candidates for novel drug target discovery;
Offer a generalizable analytical framework for systems-level studies of other rare cancers.
These advances may accelerate the translation of computational predictions into clinical applications, advancing precision oncology for rare endocrine malignancies.
Full article available on ScienceDirect:
https://doi.org/10.1016/j.intonc.2026.100056
Contact Information for Intelligent Oncology:
LinkedIn: @IntelligentOncology
X: @IntelligentOnco
Facebook: @intelligentoncology
Email Address: editorialoffice@intelligent-oncology.net
Official Website: https://www.sciencedirect.com/journal/intelligent-oncology
Submission Link: https://www2.cloud.editorialmanager.com/intonc/default2.aspx
